Research Projects
-
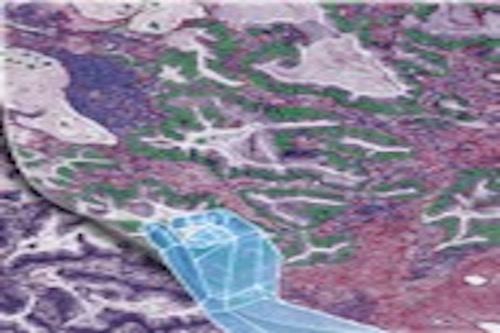

Tissue imaging data analysis
Currently, my research is mainly focused on tissue image analysis. Our team was among the first to develop and validate computational models using tissue images collected from routine clinical procedures to refine lung cancer prognosis (Luo et al, 2017, Luo et al, 2019). We have developed deep-learning-based models to detect tumor regions and micro-blood-vessels and to predict patient outcomes (Wang et al, 2018; Yi et al, 2018; Huang et al, 2017). Recently, we developed algorithms to detect and classify different types of cells from tissue images (Wang et al, 2019; Wang et al, 2020), and developed a set of spatial models (Li et al, 2019, Li et al, 2020) to investigate cell spatial organization and its implications in disease. I am the PI of a CPRIT (Cancer Prevention Institute of Texas) grant focusing on analyzing digital pathology data to improve lung cancer patient care. Our study was highlighted on the cover page of the May 2020 issue of Cancer Research. https://aacrjournals.org/cancerres/issue/80/10
- Yi F, Huang J, Yang L, Xie Y, Xiao G. Automatic extraction of cell nuclei from H&E-stained histopathological images. Journal of Medical Imaging. 2017 Apr;4(2):027502. PMID: 28653017
- Yi F, Yang L, Wang S, Lei G, Huang C; Xie Y, Xiao G*. Micro-vessel Prediction in H&E Stained Pathology Images using Fully Convolutional Neural Networks. BMC Bioinformatics 2018, 19:64. PMC5828328.
- Wang S, Wang T, Yang L, Yang DM, Fujimoto J, Yi F, Luo X, Yang Y, Yao B, Lin S, Moran C, Kalhor N, Weissferdt A, Minna J, Xie Y, Wistuba II, Mao Y, Xiao G. ConvPath: A Software Tool for Lung Adenocarcinoma Digital Pathological Image Analysis Aided by Convolutional Neural Network. EBioMedicine. 2019, 50:103-110. PMID: 31767541.
- Wang S, Rong R, Yang DM, Fujimoto J, Yan S, Cai L, Yang L, Luo D, Behrens C, Parra ER, Yao B, Xu L, Wang T, Wistuba, II, Minna J, Xie Y, Xiao G*. Computational staining of pathology images to study the tumor microenvironment in lung cancer. Cancer Research. 2020. PubMed PMID: 31915129.
-

Machine learning and deep learning
Our team has developed and validated machine learning algorithms to solve practical biological and clinical problems: (1). In 2007, I applied a machine learning method to identify new blood protein biomarkers that picks up very early stages of Alzheimer’s Disease (AD) at 88 to 96 percent accuracy with much lower cost. This blood-based test is current being validated in several large NIH-funded ongoing clinical trials as a front-line screening test for AD. (2) We developed a new algorithm to identify a gene signature to predict lung cancer patient response to adjuvant chemotherapy (Tang et al, 2013), and validate the signature using a Clinical Laboratory Improvement Amendments (CLIA)-grade assay in an independent cohort (Xie et al, 2018). (3) We developed a machine learning algorithm to predict lung cancer patient prognosis using commonly available pathology images (Luo et al, 2017), and validated the model in multiple independent patient cohorts (Luo et al, 2019).
- Xiao G, Ma S, Minna J, Xie Y. Adaptive prediction model in prospective molecular-signature-based clinical studies, Clin. Cancer Res. 2014, Feb 1;20(3):531-9. PMC3946561.
- Luo X, Zang X, Yang L, Huang J, Liang F, Rodriguez Canales J, Wistuba, II, Gazdar A, Xie Y, Xiao G*. Comprehensive Computational Pathological Image Analysis Predicts Lung Cancer Prognosis. J. Thoracic Oncol. 2017, 12:3, 501–509. PMC5462113.
- Huang C, Zhang A, Xiao G*. Deep Integrative Analysis for Survival Prediction. Pac Symp Biocomput. 2018; 23:343-352. PMID: 29218895.
- Wang S, Yang DM, Rong R, Zhan X, Xiao G*. Pathology image analysis using segmentation deep learning algorithms. American Journal of Pathology. 2019, Jun 11. PMC6723214.
-

Developing computational algorithms and bioinformatics tools for complex biomedical data
We are actively developing new bioinformatics tools and computational algorithms for big data, such as genome-wide RNAi screening data and next-generation sequencing data. My group is experienced with software development. We have developed several comprehensive web portals, including Lung Cancer Explorer (LCE) and Genomic Regression Analysis of Coordinated Expression (GRACE) (Cai, et al., Nat Communications, 2017), deep learning-based software (ConvPath), online clinical outcome prediction calculators, Galaxy-based software tools (such as PipeCLIP), and R packages. Some of this software can be accessed on my lab website: https://qbrc.swmed.edu/x-software/
- Wang T, Xie Y, Xiao G*. dCLIP: a computational approach for comparative CLIP-seq analyses. Genome Biology. 2014, Jan 7;15(1):R11.7. PMC4054096.
- Cai L, Li Q, Du Y, Yun J, Xie Y, DeBerardinis R, Xiao G*. Genomic Regression Analysis of Coordinated Expression, Nature Communications. 2017, Dec 19;8(1):2187. PMC5736603.
- Zhang M, Li Q, Yu D, Yao B, Guo W, Xie Y, Xiao G*. GeNeCK: a web server for gene network construction and visualization. BMC Bioinformatics. 2019 Jan 7;20(1):12. PMID: 30616521.
- Zhang M, Sheffield T, Zhan X, Li Q, Yang DM, Wang Y, Wang S, Xie Y, Wang T, Xiao G*. Spatial molecular profiling: platforms, applications and analysis tools. Briefings in Bioinformatics. 2020; In Press.
-
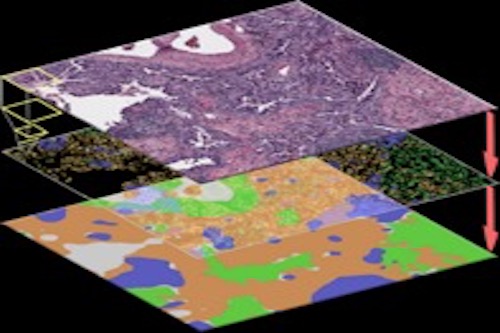

Developing spatial models for biological data
We have developed computational methodologies for Bayesian analysis, spatial modeling, and integrative analysis of different biological datasets, especially pathological imaging data.
- Zhong R, Kim M, White M, Yang X, Xiao G*. Spatial background noise correction for high-throughput RNAi screening. Bioinformatics 2013, 29(17):2218-20. PMC3740628.
- Yu D, Won S, Lim J, Xiao G*. Statistical completion of a partially identified graph with applications for the estimation of gene regulatory networks. Biostatistics, 2015, doi:10.1093/biostatistics/kxv013. PMC4570579.
- Li Q, Wang X, Liang F, Yi F, Xie Y, Gazdar A, Xiao G*. A Bayesian hidden Potts mixture model for analyzing lung cancer pathology images. Biostatistics 2018, May 18. doi: 10.1093/biostatistics/kxy019. PMC6797059.
- Li Q, Wang X, Liang F, Xiao G*, A Bayesian mark interaction model for analysis of tumor pathology images, The Annals of Applied Statistics, 2019 13 (3), 1708-1732.
-
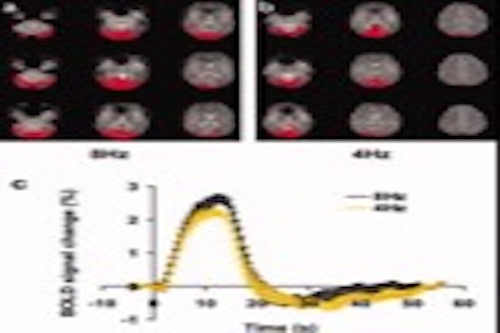
Biomedical imaging analysis
Our team has a longstanding interest and experience in biomedical image analysis. We have developed new methods and performance analysis in brain images, such as fMRI and DTI images.
- Lu H, Yezhuvath US, Xiao G. Improving fMRI sensitivity by normalization of basal physiologic state. Hum Brain Mapp. 2010;31(1):80-7. PubMed PMID: 19585589; PMCID: PMC2797559.
- Aslan S, Huang H, Uh J, Mishra V, Xiao G, van Osch MJ, Lu H. White matter cerebral blood flow is inversely correlated with structural and functional connectivity in the human brain. Neuroimage. 2011;56(3):1145-53. PubMed PMID: 21385618; PMCID: PMC3085605.
- Tung KC, Uh J, Mao D, Xu F, Xiao G, Lu H. Alterations in resting functional connectivity due to recent motor task. Neuroimage. 2013;78:316-24. PubMed PMID: 23583747; PMCID: PMC3672369.
- Liu P, Dimitrov I, Andrews T, Crane DE, Dariotis JK, Desmond J, Dumas J, Gilbert G, Kumar A, Maclntosh BJ, Tucholka A, Yang S, Xiao G, Lu H. Multisite evaluations of a T2 -relaxation-under-spin-tagging (TRUST) MRI technique to measure brain oxygenation. Magn Reson Med. 2016;75(2):680-7. PubMed PMID: 25845468; PMCID: PMC4592780.